YEL035C
Share your knowledge...Edit this entry!
| Systematic name | YEL035C |
| Gene name | UTR5 |
| Aliases | |
| Feature type | ORF, Uncharacterized |
| Coordinates | Chr V:85545..85045 |
| Primary SGDID | S000000761 |
Description of YEL035C: Protein of unknown function; transcription may be regulated by Gcr1p; essential for growth under standard (aerobic) conditions but not under anaerobic conditions[1][2]
Contents
[hide]Community Commentary
About Community Commentary. Please share your knowledge!
This gene is part of the UW-Stout Orphan Gene Project. Learn more here.
UW-Stout/Sensitivity To Nitrogen Starvation
Knocking out the YGL035C seems to have no effect on growth after incubating for 5 days on the Nitrogen omitted media.
Ultraviolet Sensitivity
YEL035C is more sensitive than BY4735(wild) given that there is less colonizes present when under the same stress.
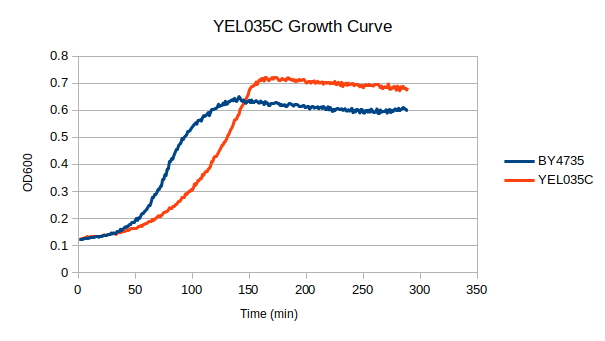
Growth Rate in YPD
In a BY4735 background, knocking out YEL035C seems to have a moderate effect on growth rate in log-phase in YPD media. In this assay, the BY4735 strain's doubling time was 124 minutes, while the YEL035C knock-out strain's doubling time was 220 minutes.
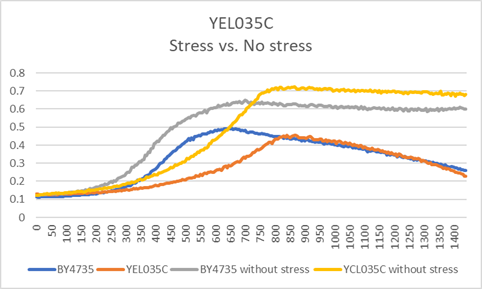
G-418 Stress
In the BY4735 background, knocking out the YEL035C strain seem to have a moderate effect on a growth rate, by slowing down the growth even more than the BY4735. In the knock-out experiment, the BY4735 strain's doubling time was 149 minutes, whereas the YEL035C knock-out strain's doubling time was 309 minutes. In the calibration experiment, the BY4735 strain's doubling time was 64 minutes, whereas the YEL035C knock-out strain's doubling time was 100 minutes.
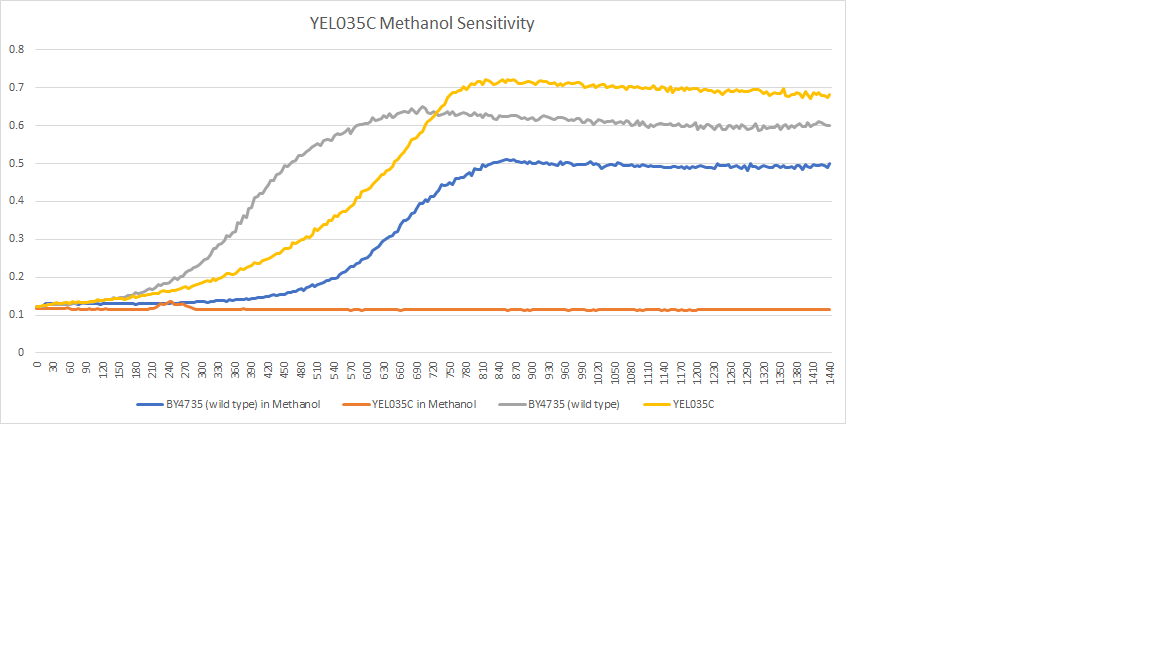
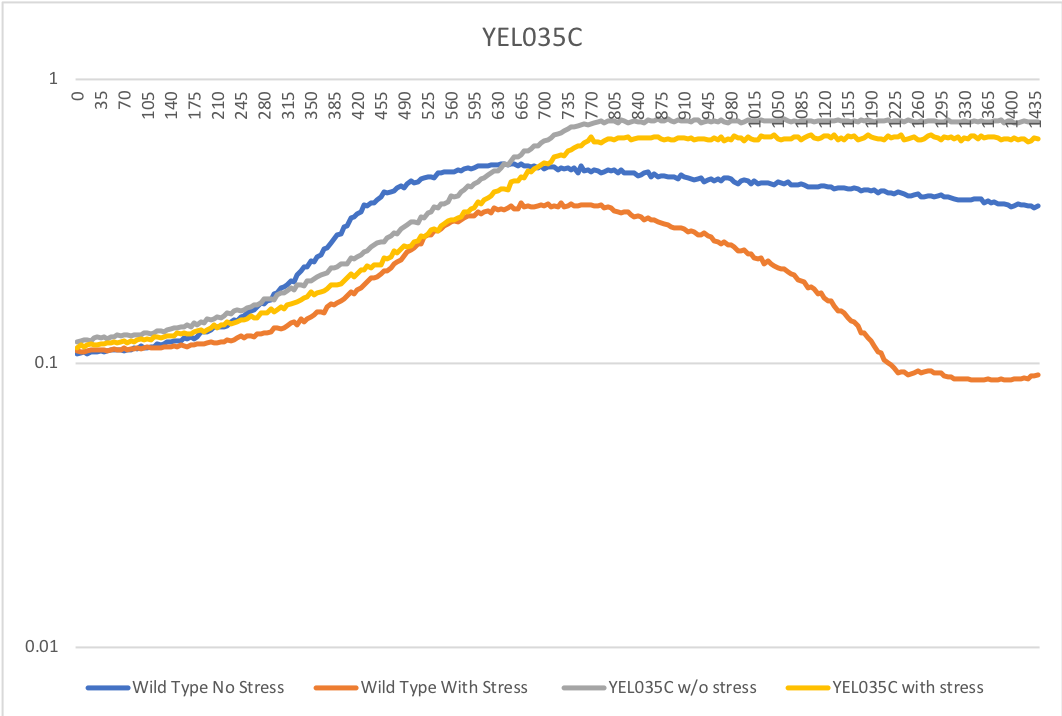
Methanol sensitivity
The wild type strain had a little difference in the doubling time when we added methanol, but the methanol killed all of the cells in the YEL035c strain. [[1]]
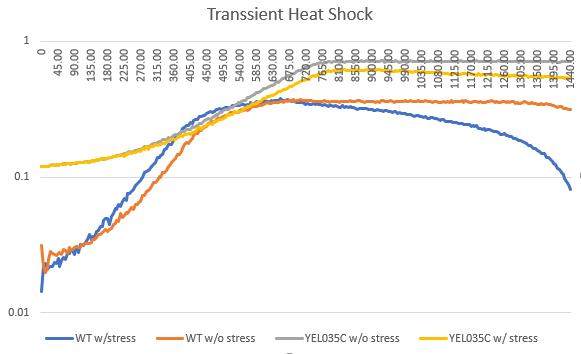
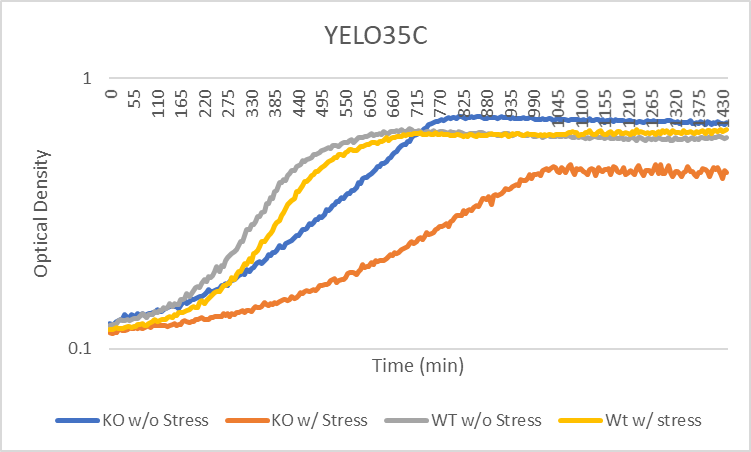
Heat Shock
Heat shock had no change on the doubling time, but the starting amount of yeast cells less. The heat seemed to kill off the cells, the longer they were in the hot water bath.
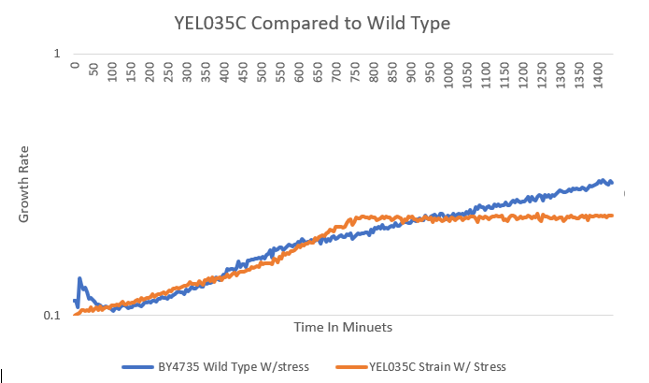
Stressing with Hydroxyurea
Hydroxyurea Protocol - The wild type strain had only a small difference in doubling time when stressed with Hydroxyurea; however, the doubling time of the YEL035C strain was much higher, indicating that the gene is more sensitive to the stress and that this gene probably plays a role in DNA replication.
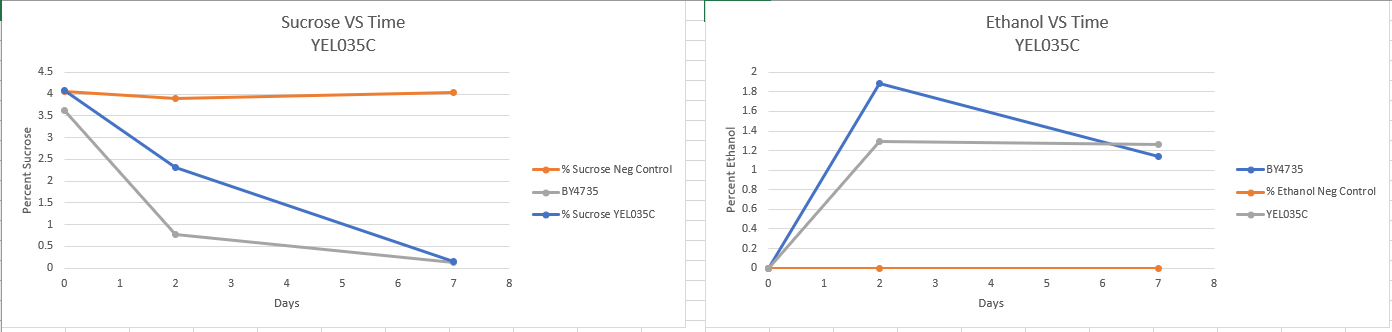
Fermentation
Fermentation protocol link [2] - There appears to be no major difference in fermentation rate. The lower final ethanol percentage could be a result of evaporation. If this experiment were to be carried out again then it should be done in an air tight container.
Caffeine Sensitivity
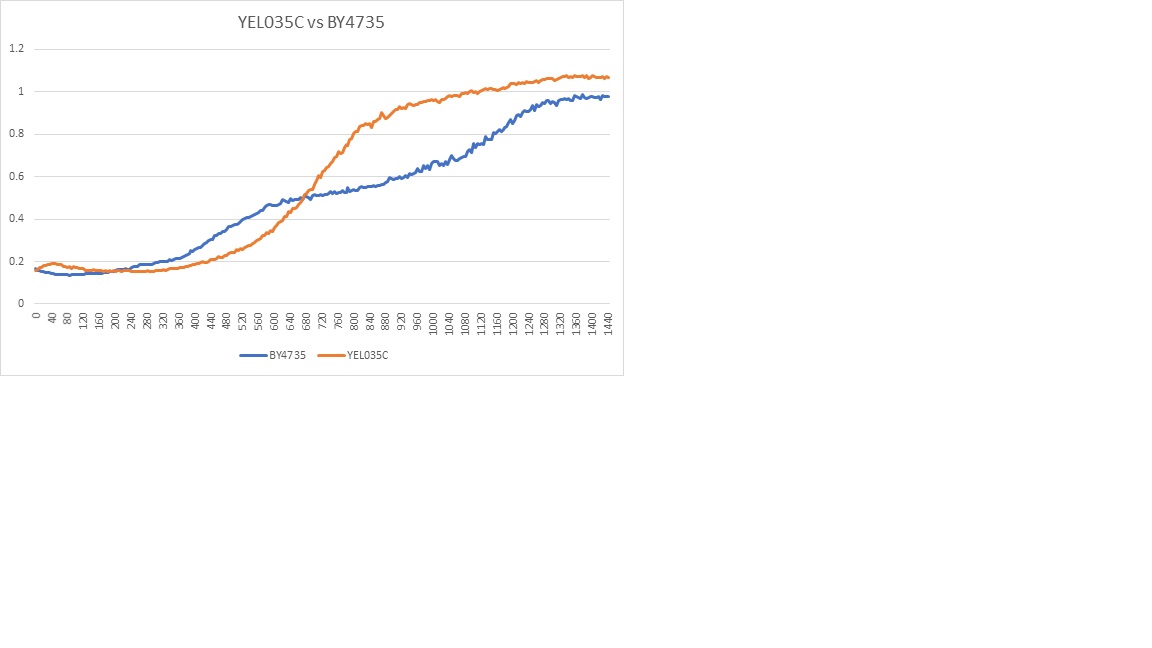 Caffeine Protocol UW-Stout/Caffeine The beginning of the graph showed the two types had very different initial reactions but became similar.
Caffeine Protocol UW-Stout/Caffeine The beginning of the graph showed the two types had very different initial reactions but became similar.
Stressing with pH
pH stress protocol link [3]
By looking at the chart here my partner and I came to the conclusion that due to the data that the chart shows when we compare the yeast strain YEL035C to the wild type strain BY4735 that the data is inconclusive to the stress test because of by looking at the chart and the way it looks like doesn’t resemble our pH stress test of 4.
Cold Shock Sensitivity
The Results from this graph have proven that compared to the wild type there was no significant change in the doubling time. This has proven that its neither more or less sensitive in comparison to the wild type.
References
See Help:References on how to add references
- Jump up ↑ Lopez MC and Baker HV (2000) Understanding the growth phenotype of the yeast gcr1 mutant in terms of global genomic expression patterns. J Bacteriol 182(17):4970-8 SGD PMID 10940042
- Jump up ↑ Snoek IS and Steensma HY (2006) Why does Kluyveromyces lactis not grow under anaerobic conditions? Comparison of essential anaerobic genes of Saccharomyces cerevisiae with the Kluyveromyces lactis genome. FEMS Yeast Res 6(3):393-403 SGD PMID 16630279
See Help:Categories on how to add the wiki page for this gene to a Category