UW-Stout/pH
Intro:
This page is dedicated to the test protocol used to view stress on yeast cells in a lowered pH environment. We started by mixing up 1 molar HCL into distilled water using a pH probe to determine when we reached the desired level. We had pH from 7 down to 4 going in roughly 1 pH step per solution. We the used these in a calibration test where we tested a unmodified wild type BY4735. This gave us a data sheet showing the growth times and effects of the varied pH levels.
Materials:
Hydrochloric Acid
Yeast Cells To test
8 Test Tubes
Proper PPE
10 ml of 1 molar ACL
Distilled water
Pipetting equipment
Ph probe
Syringe and sterilizing filter for purification.
Protocol:
1) Gather Test tubes, DI water, 1 Molar HCL, and pH probe.
2) Using the pH prob, slowly dispense HCL into a beaker of DI water while stirring the solution until the desired pH has been reached. (For this lab we used pH levels of roughly 4,5,6,7)
3) Then using these solutions, mix with your wild type cells, in a test plate. We did three wells per pH. This data is going to be your calibration to tell which pH has the most effect on the wild type.
For this lab as you can see in the data table the only pH wich really killed off the cells was in well 12, which contained pH of 4. After seeing this data, we decided to use the pH 4 solution for our strain test. Here is the calibration data:
Raw Calibration Data:
Protocol for strain test.
1) Using a sterilizing filter on a syringe, sterilize a few test tubes of solution (pH 4). Just so that you can be sure that the only growth in your plate is from wanted cells.
2)Take your sterilized water, obtain a test plate and fill 9 wells with solution (anywhere from 50 to 80 micro liters).
3)Then in each of the wells deposit a small amount of each cell type that you are testing, (roughly 50 to 30 micro liters, so that the well is at 100 micro liters)
4) in this lab we ussed, one wild type for the control, and then 8 different knock out strains for data.
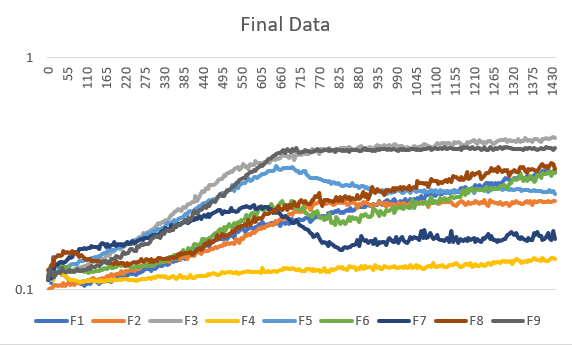
Final data:
When looking at this mess of a graph we must find the useful bits, which to us is going to be the doubling times for each line. In order to find doubling time we must use a doubling time equation which is as follows: Td=(Tf-Ti)*(((log(2))/(log((q2)/(q1))). Below is a table of all of the strands doubling time and strand name in relation to their graph labels. For in depth look at each gene and the reaction to individual data compared to the wild type, check the individual gene pages linked below.
| Gene | Doubling Time (in Min) |
|---|---|
| F1:BY4735 | Td:560 |
| F2:YEL035L | Td:483.73 |
| F3:YEL002 | Td:267.824 |
| F4:YFL064C | Td:3718.50 |
| F5:YG1235W | Td:388.57 |
| F6:YCR100C | Td:386.59 |
| F7:YFL043W | Td:511.077 |
| F8:YPL068 | Td:390.84 |
| F9:YMR090W | Td:251.55 |
Links to individual gene pages
YCL002C:[[1]]
YCR100C:[[2]]
YEL035C:[[3]]
YFL064C:[[4]]
YGL235W:[[5]]
YJL043W:[[6]]
YMR090W:[[7]]
YPL068C:[[8]]