Difference between revisions of "YNL058C"
SGDwikiBot (talk | contribs) (Automated import of articles) |
(→UW-Stout/Formamide SP22) |
||
| (40 intermediate revisions by 3 users not shown) | |||
| Line 1: | Line 1: | ||
<!-- PLEASE DO NOT EDIT HERE. USE THE SECTION EDIT LINKS ON THE RIGHT MARGIN--> | <!-- PLEASE DO NOT EDIT HERE. USE THE SECTION EDIT LINKS ON THE RIGHT MARGIN--> | ||
{{PageTop}} | {{PageTop}} | ||
| + | <protect> | ||
{|{{Prettytable}} align = 'right' width = '200px' | {|{{Prettytable}} align = 'right' width = '200px' | ||
|- | |- | ||
| − | |valign="top" nowrap bgcolor="{{SGDblue}}"| '''Systematic name''' || [http:// | + | |valign="top" nowrap bgcolor="{{SGDblue}}"| '''Systematic name''' || [http://www.yeastgenome.org/cgi-bin/locus.pl?dbid=S000005003 YNL058C] |
|- | |- | ||
|valign="top" nowrap bgcolor="{{SGDblue}}"| '''Gene name''' ||'' '' | |valign="top" nowrap bgcolor="{{SGDblue}}"| '''Gene name''' ||'' '' | ||
| Line 12: | Line 13: | ||
|- | |- | ||
|valign="top" nowrap bgcolor="{{SGDblue}}"| '''Coordinates''' | |valign="top" nowrap bgcolor="{{SGDblue}}"| '''Coordinates''' | ||
| − | |nowrap| Chr XIV: | + | |nowrap| Chr XIV:516713..515763 |
|- | |- | ||
| − | | | + | |valign="top" nowrap bgcolor="{{SGDblue}}"| '''Primary SGDID''' || S000005003 |
|} | |} | ||
<br> | <br> | ||
| − | '''Description of | + | '''Description of YNL058C:''' Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; YNL058C is not an essential gene<ref name='S000071347'>Giaever G, et al. (2002) Functional profiling of the Saccharomyces cerevisiae genome. Nature 418(6896):387-91 {{SGDpaper|S000071347}} PMID 12140549</ref><ref name='S000074185'>Huh WK, et al. (2003) Global analysis of protein localization in budding yeast. Nature 425(6959):686-91 {{SGDpaper|S000074185}} PMID 14562095</ref><ref name='S000074306'>Ubersax JA, et al. (2003) Targets of the cyclin-dependent kinase Cdk1. Nature 425(6960):859-64 |
| − | {{SGDpaper| | + | {{SGDpaper|S000074306}} PMID 14574415</ref> |
<br> | <br> | ||
<br> | <br> | ||
<br> | <br> | ||
<br> | <br> | ||
| − | + | <br> | |
| − | + | </protect> | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
__TOC__ | __TOC__ | ||
==Community Commentary== | ==Community Commentary== | ||
{{CommentaryHelp}} | {{CommentaryHelp}} | ||
| + | ===[[UW Stout/D2O SP22]]=== | ||
| + | [[Image:4.28.2022 1.png]] | ||
| + | |||
| + | The Knock Out Yeast Strain YNL058C (A6) was unaffected by the 35% dilution of D2O. This also shows in the optical density results being about 0.4-0.5 OD 600. | ||
| + | |||
| + | ===[[UW Stout/Sucrose Fermentation SP22]]=== | ||
| + | |||
| + | {| border="1" cellspacing="4" cellpadding="2" | ||
| + | |Gene | ||
| + | |Glucose | ||
| + | |Fructose | ||
| + | |Ethanol | ||
| + | |- | ||
| + | |Standard solution | ||
| + | |2.0000 | ||
| + | |0.2000 | ||
| + | |2.0000 | ||
| + | |- | ||
| + | |YDl109C | ||
| + | |0.3800 | ||
| + | |0.3933 | ||
| + | |0.3430 | ||
| + | |- | ||
| + | |YGL140C | ||
| + | |0.2212 | ||
| + | |0.2685 | ||
| + | |0.1867 | ||
| + | |- | ||
| + | |YOR111W | ||
| + | |0.3332 | ||
| + | |0.3598 | ||
| + | |0.1343 | ||
| + | |- | ||
| + | |YHL029C | ||
| + | |0.3870 | ||
| + | |0.2368 | ||
| + | |0.1151 | ||
| + | |- | ||
| + | |YDR307W | ||
| + | |0.4366 | ||
| + | |0.2487 | ||
| + | |0.0606 | ||
| + | |- | ||
| + | |YNL058C | ||
| + | |0.2710 | ||
| + | |0.3056 | ||
| + | |0.1577 | ||
| + | |- | ||
| + | |YCL049C | ||
| + | |0.4078 | ||
| + | |0.3052 | ||
| + | |0.1969 | ||
| + | |- | ||
| + | |YGR079W | ||
| + | |0.4042 | ||
| + | |0.1589 | ||
| + | |0.0080 | ||
| + | |- | ||
| + | |YBL113C | ||
| + | |0.3498 | ||
| + | |0.2012 | ||
| + | |0.1434 | ||
| + | |- | ||
| + | |BY4735 | ||
| + | |0.3171 | ||
| + | |0.3084 | ||
| + | |0.3541 | ||
| + | |} | ||
| + | |||
| + | ====Interpretation==== | ||
| + | This gene produced 44.54% of the amount of ethanol that the wild type produced. | ||
| + | |||
| + | ===[[UW-Stout/UV Light SP22]]=== | ||
| + | |||
| + | As part of the University of Wisconsin Stout [[UW-STOUT/About|Orphan Gene Project]] this gene was tested under a UV Light [[UW-Stout/UV Light SP22|using this protocol.]] | ||
| + | |||
| + | '''RESULTS''' | ||
| + | |||
| + | [[File:ASHTONMIKOLOSKIFinal Project Graph.png|450px|]] | ||
| + | [[File:AshtonGene6.jpg|350px|]] | ||
| + | |||
| + | |||
| + | '''INTERPERTATION''' | ||
| + | In the graph and photos above, exposing this gene to 600 seconds of 400 Watt UV Light killed approximately 85% of yeast cell cultures, compared to its control counterpart, which was the same gene and amount of cells, just was not exposed to UV Light. | ||
| + | |||
| + | ===[[UW-Stout/Heat Shock SP22]]=== | ||
| + | |||
| + | As part of the University of Wisconsin Stout [[UW-Stout/About|Orphan Gene Project]] this gene was tested by exposing the cells to heat shock. | ||
| + | |||
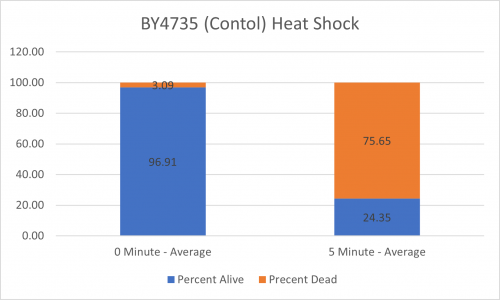
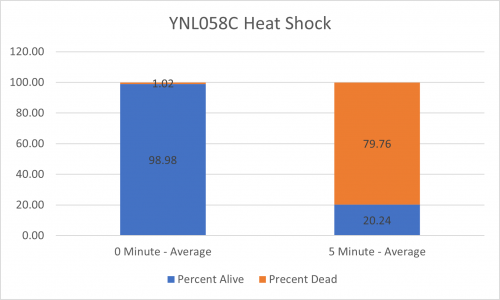
| + | ====Results==== | ||
| + | [[Image:BY4735 Control.png|BY4735 Control]] [[Image:YNL058C.png|YNL058C]] | ||
| + | [[Image:BY4735 Control Graph.png|BY4735 Control Graph|500px]] [[Image:YNL058C Graph.png|YNL058C Graph|500px]] | ||
| + | |||
| + | ====Interpretation==== | ||
| + | |||
| + | From the data gathered, there was a small negative effect to knocking out this gene when it came to the yeast's ability to hold up to heat shock. The modified yeast cells were only 83% as resistant to heat shock than the control. | ||
| + | |||
| + | ===[[UW-Stout/Caffeine SP22]]=== | ||
| + | |||
| + | As part of the University of Wisconsin Stout [[UW-Stout/About|Orphan Gene Project]] this gene was tested by exposing the cells to 4mM of caffeine. | ||
| + | |||
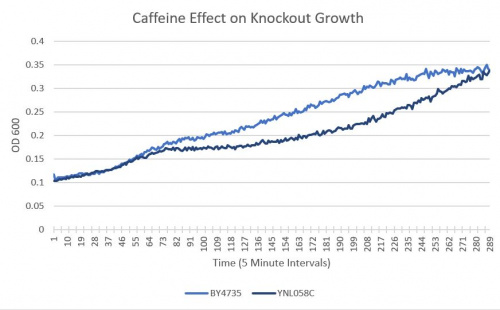
| + | ====Results==== | ||
| + | :BY4735(wild type yeast) and YNL058C(knockout yeast gene) growth after being subjected to 4mM caffeine solution [[UW-Stout/Caffeine SP22|following this protocol]]. | ||
| + | |||
| + | [[Image:YNL058C.jpeg|500px]] | ||
| + | |||
| + | ===Interpretation=== | ||
| + | As seen in the growth curve, YNL058C shows no significant difference in growth sensitivity to caffeine. It is of note that from measurement 80 to measurement 250 there is a significant difference. | ||
| + | |||
| + | ====Interpretation==== | ||
| + | |||
| + | |||
| + | ===[[UW-Stout/Hydrogen Peroxide SP22]]=== | ||
| + | |||
| + | As part of the University of Wisconsin Stout Orphan Gene Project this gene was tested by exposing the cells to hydrogen peroxide. | ||
| + | |||
| + | |||
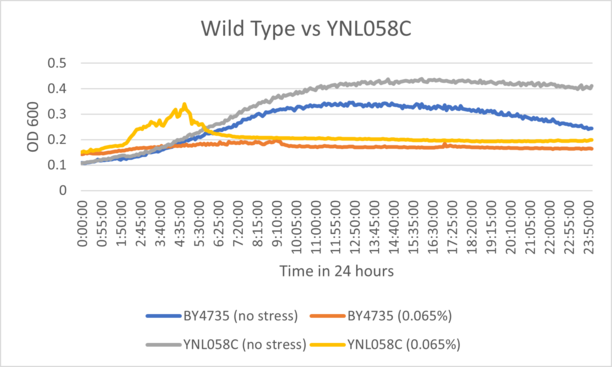
| + | ====Results==== | ||
| + | |||
| + | BY4735 and YNL058C after being exposed to 0.065% dilution of hydrogen peroxide solution. | ||
| + | |||
| + | [[File:YNL058Cperosm.png]] | ||
| + | |||
| + | ====Interpretation==== | ||
| + | The growth curve for YNL058C suggests that the knockout had very little variation from wild type cells when exposed to hydrogen peroxide. The growth ceiling for the unstressed control was higher than wild type after 24 hours, suggesting the strain may have fast initial growth. | ||
| + | ===[[UW Stout/Nystatin SP22]]=== | ||
| + | As part of the University of Wisconsin Stout [[UW-Stout/About|Orphan Gene Project]] this gene was tested by exposing the cells to 1µg/ml Nystatin solution. | ||
| + | ====Results==== | ||
| + | [[File:YNL-T1.jpg]] | ||
| + | [[File:YNL-T2.jpg]] | ||
| + | |||
| + | |||
| + | Based on the growth curves, we can see that the YNL058C control is growing at a faster rate than both the wild type control and the stressed wild type. However, the YNL058C being treated with Nystatin is growing at a slower rate than both wild type control and stressed wild type. This indicates that the removal of the YNL058C gene lessens the cells resistance to Nystatin. | ||
| + | |||
| + | |||
| + | ===[[UW-Stout/Formamide SP22]]=== | ||
| + | |||
| + | ===Results=== | ||
| + | [[File:wildyeast.jpg|400px|]] [[File:yeast.jpg|400px|]] [[File:doublingtimestable.jpg|400px|]] | ||
| + | |||
| + | |||
| + | *3% formamide line graph: x-axis= growing time (min); y-axis= optical density 600 | ||
| + | *Trial 2 line graph: x-axis= growing time (min); y-axis= optical density 600; key= yeast strains and corresponding color to the individual lines in the line graph | ||
| + | *Average doubling time bar graph: x-axis= yeast strains and corresponding colors to the trial 2 line graph; y-axis= average doubling time (min) | ||
| + | |||
| + | ===Interpretation=== | ||
| + | The wild yeast cell treated with 3% formamide had a doubling time of 876.69 min. The 3% formamide solution added to nine transformed yeast cells was then compared to the wild yeast cell using the computed doubling times to see the effects of the formamide. In the experiment, some of the transformed yeast cells were heavily effected by the formamide, however, the YNL058C strain was one of the few not heavily effected- in comparison with the wild yeast cells. The YNL058C strain had doubling times slightly faster than the wild yeast cell, with a doubling time of 844.12 min. It can be concluded that the transformed yeast cell, YNL058C, did not have a unique response to the formamide stress test used in the experiment. | ||
| + | |||
| + | == Transmembrane Domain GxxxG == | ||
| + | |||
| + | The transmembrane domain of Ynl058cp shows homology with the SKG6/AXL2 alpha-helix transmembrane domain (IPR014805). In particular, this domain contains GaviG. The presence of a overlapping GxxxG motif <ref name='PMID 15313242'> Senes A, Engel DE and DeGrado WF (2004) Folding of helical membrane proteins: the role of polar, GxxxG-like and proline motifs. Curr Opin Struct Biol. 2004 14(4):465-79. PMID 15313242</ref> strongly indicates that Ynl058c dimerises. | ||
| + | |||
=== Protein Details === | === Protein Details === | ||
[[Category:Topic:Protein Details]] | [[Category:Topic:Protein Details]] | ||
| Line 44: | Line 190: | ||
{{ShortCenteredHR}} | {{ShortCenteredHR}} | ||
| + | |||
| + | {{UW-Stout}} | ||
| + | |||
| + | <!-- PLEASE ADD Community Commentary ABOVE THIS MESSAGE. See below for an example of community annotation --> | ||
| + | <!-- | ||
| + | Specifically higher expression in carbon limited chemostat cultures versus carbon excess. | ||
| + | <ref>Boer VM, et al. (2003) The genome-wide transcriptional responses of Saccharomyces cerevisiae grown on glucose in aerobic chemostat cultures limited for carbon, nitrogen, phosphorus, or sulfur. | ||
| + | J Biol Chem 278(5):3265-74</ref> | ||
| + | --> | ||
| + | |||
| + | |||
| + | |||
| + | <protect> | ||
==References== | ==References== | ||
<!-- REFERENCES ARE AUTOMATICALLY GENERATED. PLEASE DON'T EDIT THIS SECTION--> | <!-- REFERENCES ARE AUTOMATICALLY GENERATED. PLEASE DON'T EDIT THIS SECTION--> | ||
{{RefHelp}} | {{RefHelp}} | ||
| + | </protect> | ||
| + | |||
| + | '''[[UW-Stout/Nonionic Detergent SP22]]''' | ||
| + | |||
| + | As part of the University of Wisconsin Stout Orphan Gene Project this gene was tested by exposing the cells to 0.001% concentrated NLS. (Nonionic detergent) | ||
| + | |||
| + | [[File:YNL058C.png|500px]] | ||
| + | |||
| + | '''Interpretations:''' | ||
| + | |||
| + | -According to the graph above, we can see that the KO yeast cell YNL058C follow a smaller sensitivity to the stress than the Wild type strain. | ||
| + | |||
| + | -We can then conclude that the knocked-out gene in YNL058C doesn't have a significant impact on the growth of the yeast cell. | ||
| + | |||
| + | |||
| + | '''[[UW-Stout/pH Acid SP22]]''' | ||
| + | |||
| + | [[File:Strain6.jpg|350px|]] | ||
| + | Knocked out gene seems to have very little effect on cell growth in pH 7 environment based on the flattening of the growth curve. Spikes in stressed line are to be ignored; they occurred due to clumping of cells during analysis. We are focusing on the general linear trend of the growth curve. Please see protocol for specific quantitative doubling time results. | ||
| + | |||
| + | |||
| + | [[File:Wild (1).jpg|350px|]] | ||
| + | The wild-type strain was used as a control in this experiment; no genes were knocked out, it just represents how the experiment ran on the knocked-out gene strains affects a "typical" cell growth curve. Inclusion of this data on each knock-out strain growth curve graph made for difficult interpretation, so it has been separated. Please refer to this graph as a control when viewing the knock-out strain growth curve graph. Spikes in stressed line are to be ignored; they occurred due to clumping of cells during analysis. We are focusing on the general linear trend of the growth curve. | ||
Latest revision as of 19:34, 11 May 2022
Share your knowledge...Edit this entry! <protect>
| Systematic name | YNL058C |
| Gene name | |
| Aliases | |
| Feature type | ORF, Uncharacterized |
| Coordinates | Chr XIV:516713..515763 |
| Primary SGDID | S000005003 |
Description of YNL058C: Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; YNL058C is not an essential gene[1][2][3]
</protect>
Contents
- 1 Community Commentary
- 2 Transmembrane Domain GxxxG
- 3 References
Community Commentary
About Community Commentary. Please share your knowledge!
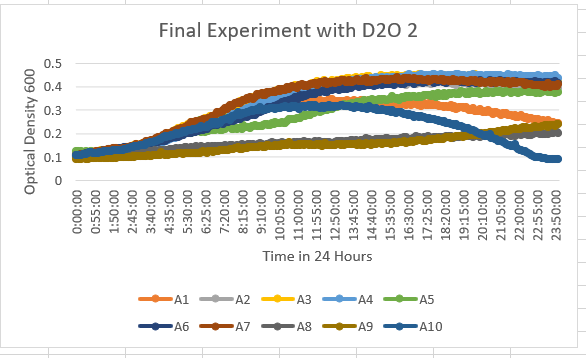
UW Stout/D2O SP22
The Knock Out Yeast Strain YNL058C (A6) was unaffected by the 35% dilution of D2O. This also shows in the optical density results being about 0.4-0.5 OD 600.
UW Stout/Sucrose Fermentation SP22
| Gene | Glucose | Fructose | Ethanol |
| Standard solution | 2.0000 | 0.2000 | 2.0000 |
| YDl109C | 0.3800 | 0.3933 | 0.3430 |
| YGL140C | 0.2212 | 0.2685 | 0.1867 |
| YOR111W | 0.3332 | 0.3598 | 0.1343 |
| YHL029C | 0.3870 | 0.2368 | 0.1151 |
| YDR307W | 0.4366 | 0.2487 | 0.0606 |
| YNL058C | 0.2710 | 0.3056 | 0.1577 |
| YCL049C | 0.4078 | 0.3052 | 0.1969 |
| YGR079W | 0.4042 | 0.1589 | 0.0080 |
| YBL113C | 0.3498 | 0.2012 | 0.1434 |
| BY4735 | 0.3171 | 0.3084 | 0.3541 |
Interpretation
This gene produced 44.54% of the amount of ethanol that the wild type produced.
UW-Stout/UV Light SP22
As part of the University of Wisconsin Stout Orphan Gene Project this gene was tested under a UV Light using this protocol.
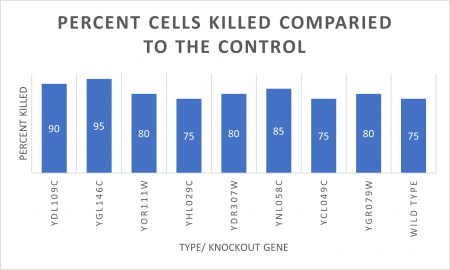
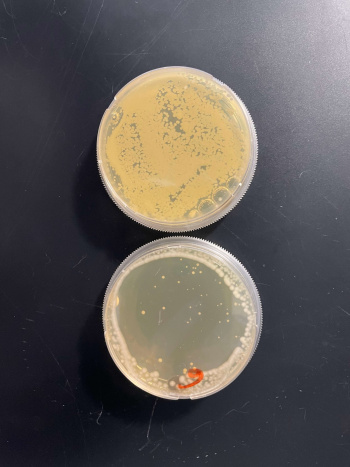
RESULTS
INTERPERTATION
In the graph and photos above, exposing this gene to 600 seconds of 400 Watt UV Light killed approximately 85% of yeast cell cultures, compared to its control counterpart, which was the same gene and amount of cells, just was not exposed to UV Light.
UW-Stout/Heat Shock SP22
As part of the University of Wisconsin Stout Orphan Gene Project this gene was tested by exposing the cells to heat shock.
Results
Interpretation
From the data gathered, there was a small negative effect to knocking out this gene when it came to the yeast's ability to hold up to heat shock. The modified yeast cells were only 83% as resistant to heat shock than the control.
UW-Stout/Caffeine SP22
As part of the University of Wisconsin Stout Orphan Gene Project this gene was tested by exposing the cells to 4mM of caffeine.
Results
- BY4735(wild type yeast) and YNL058C(knockout yeast gene) growth after being subjected to 4mM caffeine solution following this protocol.
Interpretation
As seen in the growth curve, YNL058C shows no significant difference in growth sensitivity to caffeine. It is of note that from measurement 80 to measurement 250 there is a significant difference.
Interpretation
UW-Stout/Hydrogen Peroxide SP22
As part of the University of Wisconsin Stout Orphan Gene Project this gene was tested by exposing the cells to hydrogen peroxide.
Results
BY4735 and YNL058C after being exposed to 0.065% dilution of hydrogen peroxide solution.
Interpretation
The growth curve for YNL058C suggests that the knockout had very little variation from wild type cells when exposed to hydrogen peroxide. The growth ceiling for the unstressed control was higher than wild type after 24 hours, suggesting the strain may have fast initial growth.
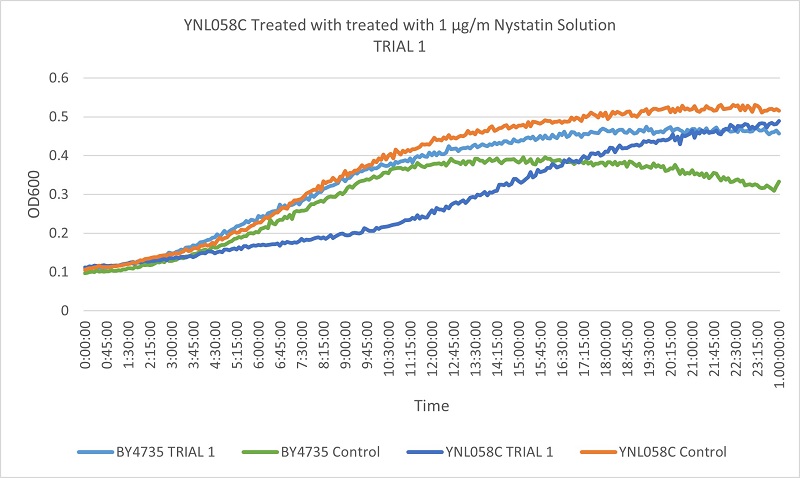
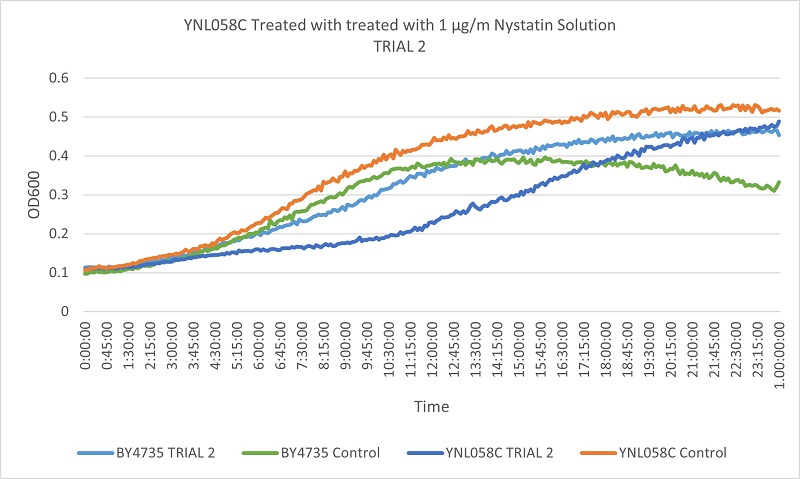
UW Stout/Nystatin SP22
As part of the University of Wisconsin Stout Orphan Gene Project this gene was tested by exposing the cells to 1µg/ml Nystatin solution.
Results
Based on the growth curves, we can see that the YNL058C control is growing at a faster rate than both the wild type control and the stressed wild type. However, the YNL058C being treated with Nystatin is growing at a slower rate than both wild type control and stressed wild type. This indicates that the removal of the YNL058C gene lessens the cells resistance to Nystatin.
UW-Stout/Formamide SP22
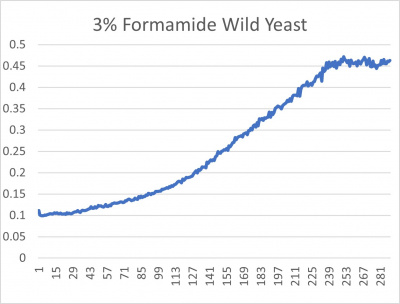
Results
- 3% formamide line graph: x-axis= growing time (min); y-axis= optical density 600
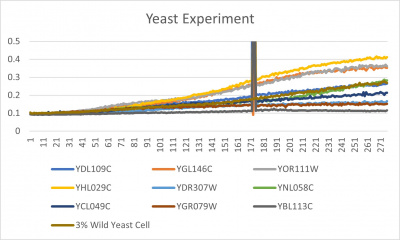
- Trial 2 line graph: x-axis= growing time (min); y-axis= optical density 600; key= yeast strains and corresponding color to the individual lines in the line graph
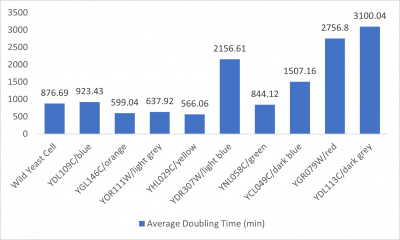
- Average doubling time bar graph: x-axis= yeast strains and corresponding colors to the trial 2 line graph; y-axis= average doubling time (min)
Interpretation
The wild yeast cell treated with 3% formamide had a doubling time of 876.69 min. The 3% formamide solution added to nine transformed yeast cells was then compared to the wild yeast cell using the computed doubling times to see the effects of the formamide. In the experiment, some of the transformed yeast cells were heavily effected by the formamide, however, the YNL058C strain was one of the few not heavily effected- in comparison with the wild yeast cells. The YNL058C strain had doubling times slightly faster than the wild yeast cell, with a doubling time of 844.12 min. It can be concluded that the transformed yeast cell, YNL058C, did not have a unique response to the formamide stress test used in the experiment.
Transmembrane Domain GxxxG
The transmembrane domain of Ynl058cp shows homology with the SKG6/AXL2 alpha-helix transmembrane domain (IPR014805). In particular, this domain contains GaviG. The presence of a overlapping GxxxG motif [4] strongly indicates that Ynl058c dimerises.
Protein Details
Protein Modification
Modification(s): Phosphorylation
Identified as an efficient substrate of Clb2-Cdk1-as1 in a screen of a proteomic GST-fusion library. [3] [5]
This gene is part of the UW-Stout Orphan Gene Project. Learn more here.
<protect>
References
See Help:References on how to add references
- ↑ Giaever G, et al. (2002) Functional profiling of the Saccharomyces cerevisiae genome. Nature 418(6896):387-91 SGD PMID 12140549
- ↑ Huh WK, et al. (2003) Global analysis of protein localization in budding yeast. Nature 425(6959):686-91 SGD PMID 14562095
- ↑ 3.0 3.1 Ubersax JA, et al. (2003) Targets of the cyclin-dependent kinase Cdk1. Nature 425(6960):859-64
SGD PMID 14574415 Cite error: Invalid
<ref>tag; name "S000074306" defined multiple times with different content - ↑ Senes A, Engel DE and DeGrado WF (2004) Folding of helical membrane proteins: the role of polar, GxxxG-like and proline motifs. Curr Opin Struct Biol. 2004 14(4):465-79. PMID 15313242
- ↑ submitted by Jeff Ubersax on 2004-01-29
See Help:Categories on how to add the wiki page for this gene to a Category </protect>
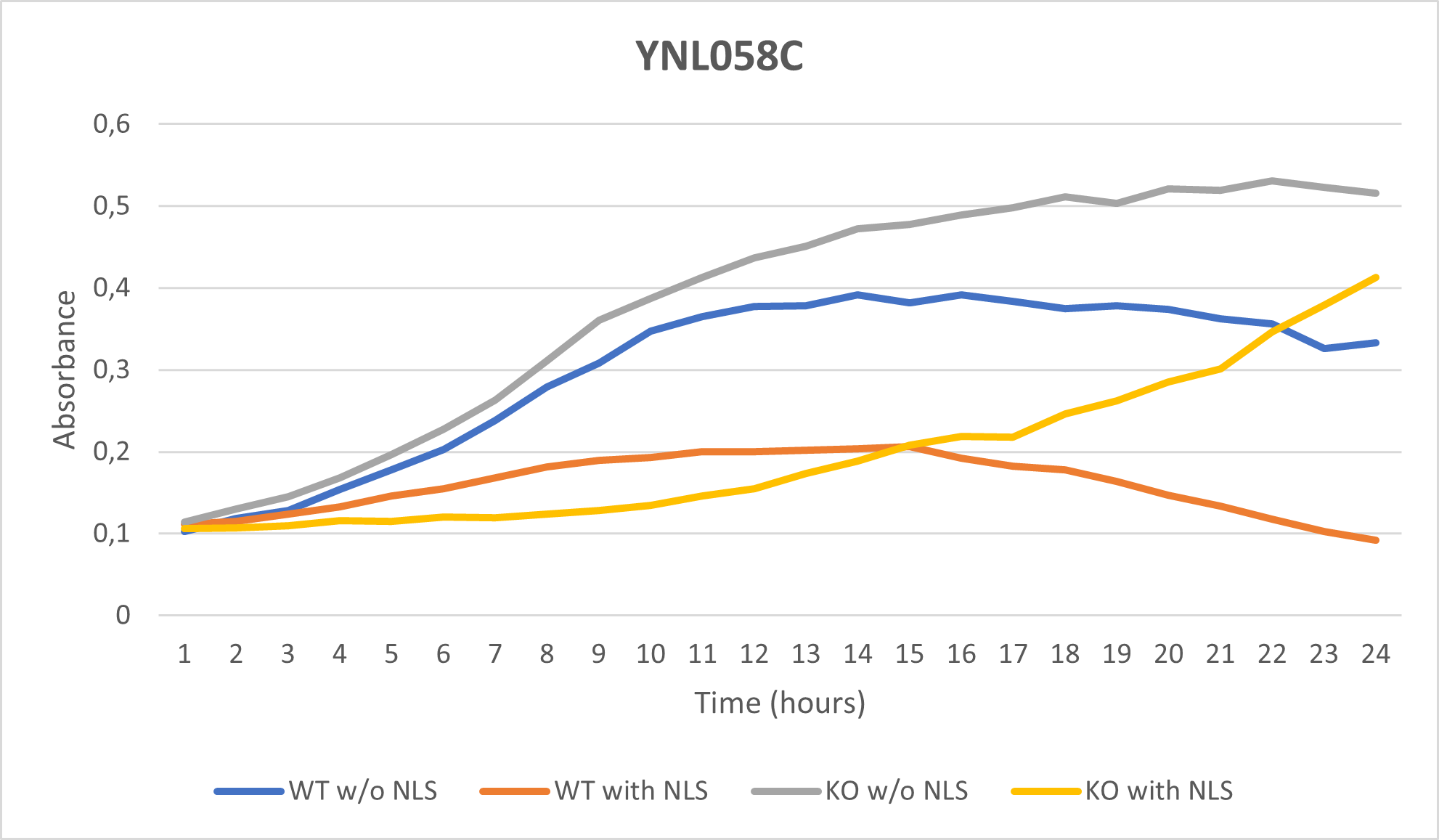
UW-Stout/Nonionic Detergent SP22
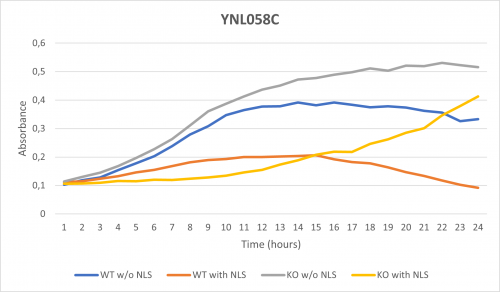
As part of the University of Wisconsin Stout Orphan Gene Project this gene was tested by exposing the cells to 0.001% concentrated NLS. (Nonionic detergent)
Interpretations:
-According to the graph above, we can see that the KO yeast cell YNL058C follow a smaller sensitivity to the stress than the Wild type strain.
-We can then conclude that the knocked-out gene in YNL058C doesn't have a significant impact on the growth of the yeast cell.
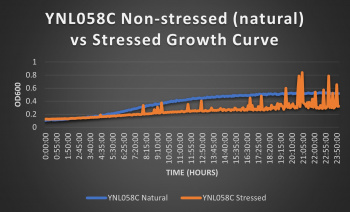 Knocked out gene seems to have very little effect on cell growth in pH 7 environment based on the flattening of the growth curve. Spikes in stressed line are to be ignored; they occurred due to clumping of cells during analysis. We are focusing on the general linear trend of the growth curve. Please see protocol for specific quantitative doubling time results.
Knocked out gene seems to have very little effect on cell growth in pH 7 environment based on the flattening of the growth curve. Spikes in stressed line are to be ignored; they occurred due to clumping of cells during analysis. We are focusing on the general linear trend of the growth curve. Please see protocol for specific quantitative doubling time results.
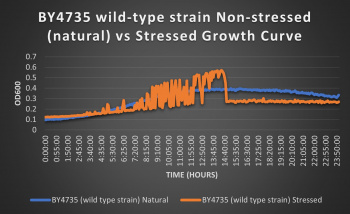 The wild-type strain was used as a control in this experiment; no genes were knocked out, it just represents how the experiment ran on the knocked-out gene strains affects a "typical" cell growth curve. Inclusion of this data on each knock-out strain growth curve graph made for difficult interpretation, so it has been separated. Please refer to this graph as a control when viewing the knock-out strain growth curve graph. Spikes in stressed line are to be ignored; they occurred due to clumping of cells during analysis. We are focusing on the general linear trend of the growth curve.
The wild-type strain was used as a control in this experiment; no genes were knocked out, it just represents how the experiment ran on the knocked-out gene strains affects a "typical" cell growth curve. Inclusion of this data on each knock-out strain growth curve graph made for difficult interpretation, so it has been separated. Please refer to this graph as a control when viewing the knock-out strain growth curve graph. Spikes in stressed line are to be ignored; they occurred due to clumping of cells during analysis. We are focusing on the general linear trend of the growth curve.