Difference between revisions of "YOR387C"
(→Hydroxychloroquine) |
m |
||
| Line 41: | Line 41: | ||
[[Image:YOR387C.png]] | [[Image:YOR387C.png]] | ||
| − | |||
| − | |||
Under normal conditions, the BY4753 doubling time is 144 minutes, while that of YOR387C is 158 minutes. On average, the YOR387C lag phase was slightly longer than that of BY4753 under normal conditions. When grown in an environment with hydroxychloroquine, YOR387C doubled in 249 minutes, while BY4753 doubled in 276 minutes. This shows that hydroxychloroquine inhibits BY4753 slightly more than YOR387C. (These doubling times and curves are the means of three experiments.) | Under normal conditions, the BY4753 doubling time is 144 minutes, while that of YOR387C is 158 minutes. On average, the YOR387C lag phase was slightly longer than that of BY4753 under normal conditions. When grown in an environment with hydroxychloroquine, YOR387C doubled in 249 minutes, while BY4753 doubled in 276 minutes. This shows that hydroxychloroquine inhibits BY4753 slightly more than YOR387C. (These doubling times and curves are the means of three experiments.) | ||
| Line 52: | Line 50: | ||
J Biol Chem 278(5):3265-74</ref> | J Biol Chem 278(5):3265-74</ref> | ||
--> | --> | ||
| + | ---- | ||
| + | ===[[UW-Stout/Salt_Concentration_SP21|Salt Concentration (NaCl)]]=== | ||
| + | |||
| + | [[Image:NaCl-BY4735vYOR387C-Graph.JPG]] | ||
| + | |||
| + | |||
| + | 0mM NaCl conc. BY4735 strain's doubling time: 169 minutes | ||
| + | |||
| + | 0mM NaCl conc. YOR387C strain's doubling time: 129 minutes | ||
| + | |||
| + | 750mM NaCl conc. BY4735 strain's doubling time: 294 minutes | ||
| + | |||
| + | 750mM NaCl conc. YOR387C strain's doubling time: 282 minutes | ||
| + | |||
| + | The graph above shows the growth rate for the previously listed strains and the relative level of NaCl concentration. Comparing the YOR387C doubling times in 0mM and 750mM NaCl, there is a large effect on the growth rate. This effect negatively impacts the knockout strain's (YOR387C) growth rate. | ||
| + | ---- | ||
Revision as of 10:58, 27 April 2021
Share your knowledge...Edit this entry! <protect>
| Systematic name | YOR387C |
| Gene name | |
| Aliases | |
| Feature type | ORF, Uncharacterized |
| Coordinates | Chr XV:1070241..1069621 |
| Primary SGDID | S000005914 |
Description of YOR387C: Putative protein of unknown function; regulated by the metal-responsive Aft1p transcription factor; highly inducible in zinc-depleted conditions; localizes to the soluble fraction[1][2][3]
</protect>
Contents
Community Commentary
About Community Commentary. Please share your knowledge!
This gene is part of the UW-Stout Orphan Gene Project. Learn more here.
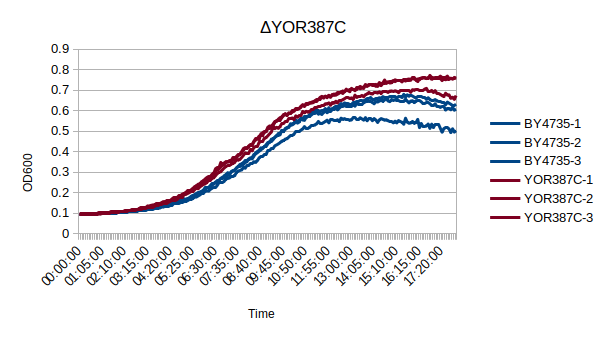
Growth Curve
In a BY4735 background, knocking out YOR387C does not seem to have much effect on the strain's growth rate. In this experiment, the BY4735 strain's doubling time was 162 minutes, while the YOR387C knock-out strain's doubling time was 137 minutes. (These doubling times are the means of three experiments.)
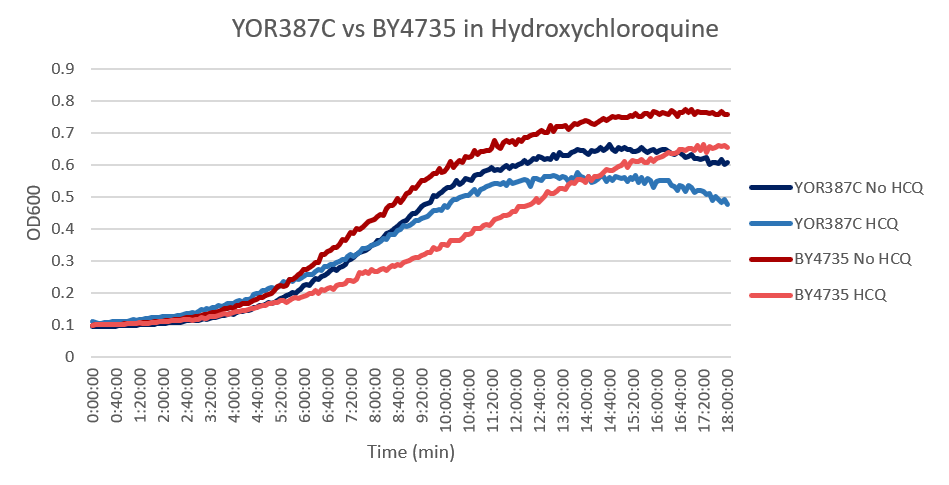
Hydroxychloroquine
Under normal conditions, the BY4753 doubling time is 144 minutes, while that of YOR387C is 158 minutes. On average, the YOR387C lag phase was slightly longer than that of BY4753 under normal conditions. When grown in an environment with hydroxychloroquine, YOR387C doubled in 249 minutes, while BY4753 doubled in 276 minutes. This shows that hydroxychloroquine inhibits BY4753 slightly more than YOR387C. (These doubling times and curves are the means of three experiments.)
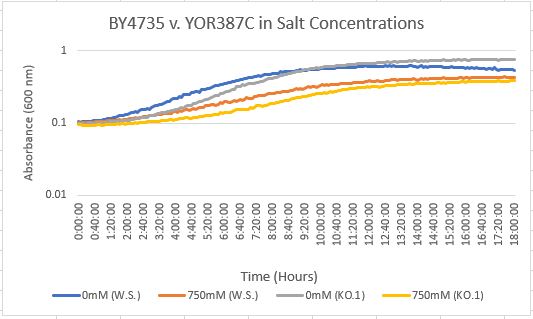
Salt Concentration (NaCl)
0mM NaCl conc. BY4735 strain's doubling time: 169 minutes
0mM NaCl conc. YOR387C strain's doubling time: 129 minutes
750mM NaCl conc. BY4735 strain's doubling time: 294 minutes
750mM NaCl conc. YOR387C strain's doubling time: 282 minutes
The graph above shows the growth rate for the previously listed strains and the relative level of NaCl concentration. Comparing the YOR387C doubling times in 0mM and 750mM NaCl, there is a large effect on the growth rate. This effect negatively impacts the knockout strain's (YOR387C) growth rate.
<protect>
References
See Help:References on how to add references
- ↑ Higgins VJ, et al. (2003) Application of genome-wide expression analysis to identify molecular markers useful in monitoring industrial fermentations. Appl Environ Microbiol 69(12):7535-40 SGD PMID 14660410
- ↑ Rutherford JC, et al. (2003) Aft1p and Aft2p mediate iron-responsive gene expression in yeast through related promoter elements. J Biol Chem 278(30):27636-43 SGD PMID 12756250
- ↑ Terashima H, et al. (2002) Sequence-based approach for identification of cell wall proteins in Saccharomyces cerevisiae. Curr Genet 40(5):311-6 SGD PMID 11935221
See Help:Categories on how to add the wiki page for this gene to a Category </protect>