Difference between revisions of "YBR284W"
(→Community Commentary) |
(→pH) |
||
| Line 44: | Line 44: | ||
[[File:pH_knockout.jpg]] | [[File:pH_knockout.jpg]] | ||
| − | + | ||
| + | YBR284w | ||
*When this strain was exposed to a citric acid buffer with a pH of 2.6, there did not appear to be a significant change in the growth rate of the strain | *When this strain was exposed to a citric acid buffer with a pH of 2.6, there did not appear to be a significant change in the growth rate of the strain | ||
Revision as of 13:10, 14 December 2021
Share your knowledge...Edit this entry! <protect>
| Systematic name | YBR284W |
| Gene name | |
| Aliases | |
| Feature type | ORF, Uncharacterized |
| Coordinates | Chr II:771239..773632 |
| Primary SGDID | S000000488 |
Description of YBR284W: Protein of unknown function; some similarity to AMP deaminases but lacks key catalytic residues and does not rescue purine nucleotide metabolic defect of quadruple aah1 ade8 amd1 his1 mutant; null mutant exhibits longer telomeres, altered Ty mobility, decreased resistance to rapamycin and wortmannin, and synthetic phenotype with expression of alpha-synuclein; induced in response to hydrostatic pressure; not an essential gene[1][2][3][4][5][6][7]
</protect>
Contents
Community Commentary
About Community Commentary. Please share your knowledge!
This gene is part of the UW-Stout Orphan Gene Project. Learn more here.
UW-Stout/Heat Shock FA21
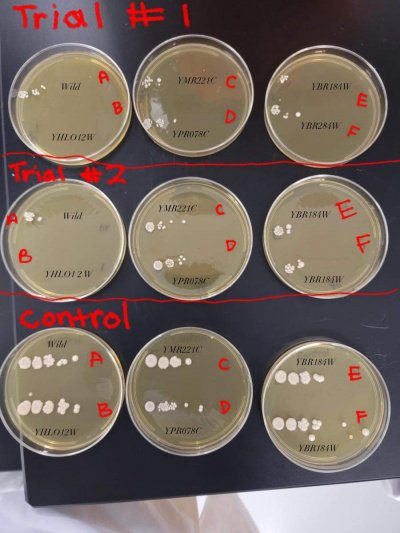
YBR284W (F)
- This yeast strain had strong growth in the control group but was very stressed by the heat shock and showed less clusters/colonies on the heat shocked trial plates. The heat shocked trial plates showed a growth decrease by 4 times.
Nitrogen Starvation
- For the YBR284W gene, results showed that there was a slight increase in growth on the dish that did not contain NH₄ where there were more colonies of yeast.
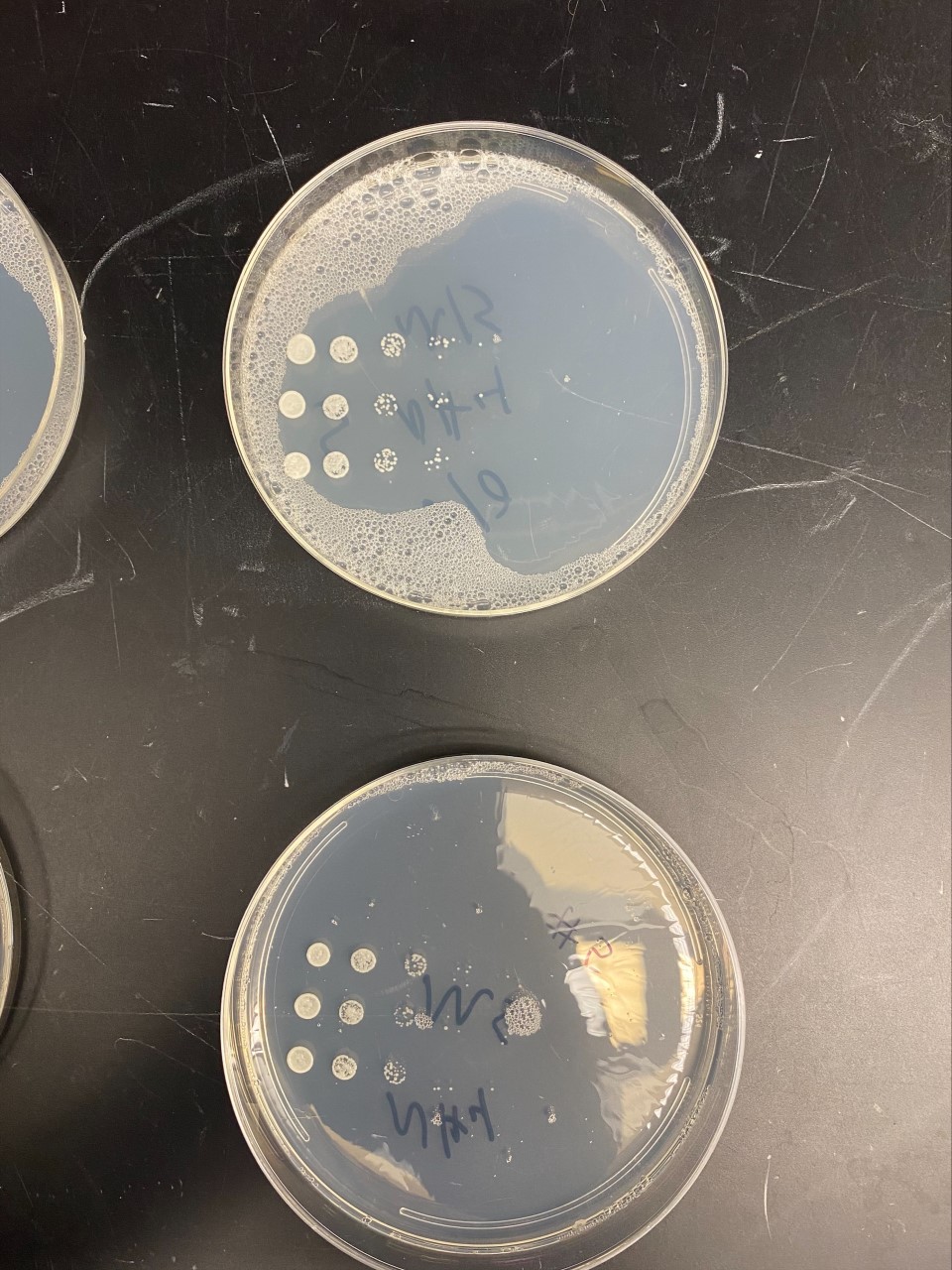
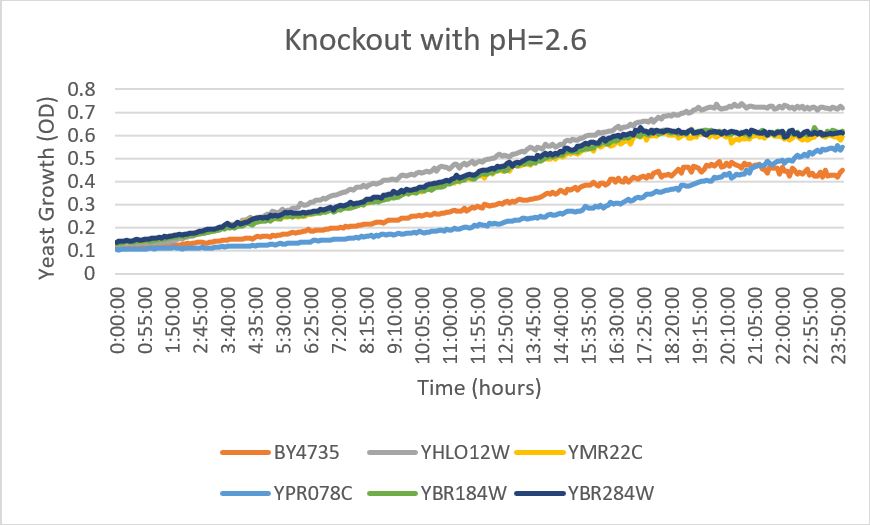
pH
YBR284w
- When this strain was exposed to a citric acid buffer with a pH of 2.6, there did not appear to be a significant change in the growth rate of the strain
<protect>
References
See Help:References on how to add references
- ↑ Gatbonton T, et al. (2006) Telomere length as a quantitative trait: genome-wide survey and genetic mapping of telomere length-control genes in yeast. PLoS Genet 2(3):e35 SGD PMID 16552446
- ↑ Holmstrom K, et al. (1994) The sequence of a 32,420 bp segment located on the right arm of chromosome II from Saccharomyces cerevisiae. Yeast 10 Suppl A:S47-62 SGD PMID 8091861
- ↑ Iwahashi H, et al. (2003) Piezophysiology of genome wide gene expression levels in the yeast Saccharomyces cerevisiae. Extremophiles 7(4):291-8 SGD PMID 12910389
- ↑ Maxwell PH and Curcio MJ (2007) Host factors that control long terminal repeat retrotransposons in Saccharomyces cerevisiae: implications for regulation of mammalian retroviruses. Eukaryot Cell 6(7):1069-80 SGD PMID 17496126
- ↑ Saint-Marc C, et al. (2009) Phenotypic consequences of purine nucleotide imbalance in Saccharomyces cerevisiae. Genetics 183(2):529-38, 1SI-7SI SGD PMID 19635936
- ↑ Willingham S, et al. (2003) Yeast genes that enhance the toxicity of a mutant huntingtin fragment or alpha-synuclein. Science 302(5651):1769-72 SGD PMID 14657499
- ↑ Xie MW, et al. (2005) Insights into TOR function and rapamycin response: chemical genomic profiling by using a high-density cell array method. Proc Natl Acad Sci U S A 102(20):7215-20 SGD PMID 15883373
See Help:Categories on how to add the wiki page for this gene to a Category </protect>