Difference between revisions of "YPR078C"
SGDwikiBot (talk | contribs) (Automated import of articles) |
(→UW-Stout/Copper FA21) |
||
| (28 intermediate revisions by 2 users not shown) | |||
| Line 4: | Line 4: | ||
{|{{Prettytable}} align = 'right' width = '200px' | {|{{Prettytable}} align = 'right' width = '200px' | ||
|- | |- | ||
| − | |valign="top" nowrap bgcolor="{{SGDblue}}"| '''Systematic name''' || [http:// | + | |valign="top" nowrap bgcolor="{{SGDblue}}"| '''Systematic name''' || [http://www.yeastgenome.org/cgi-bin/locus.pl?dbid=S000006282 YPR078C] |
|- | |- | ||
|valign="top" nowrap bgcolor="{{SGDblue}}"| '''Gene name''' ||'' '' | |valign="top" nowrap bgcolor="{{SGDblue}}"| '''Gene name''' ||'' '' | ||
| Line 13: | Line 13: | ||
|- | |- | ||
|valign="top" nowrap bgcolor="{{SGDblue}}"| '''Coordinates''' | |valign="top" nowrap bgcolor="{{SGDblue}}"| '''Coordinates''' | ||
| − | |nowrap| Chr XVI: | + | |nowrap| Chr XVI:698265..697147 |
|- | |- | ||
|valign="top" nowrap bgcolor="{{SGDblue}}"| '''Primary SGDID''' || S000006282 | |valign="top" nowrap bgcolor="{{SGDblue}}"| '''Primary SGDID''' || S000006282 | ||
|} | |} | ||
<br> | <br> | ||
| − | '''Description of YPR078C:''' Putative protein of unknown function; possible role in DNA metabolism and/or in genome stability; expression is heat-inducible<ref name=' | + | '''Description of YPR078C:''' Putative protein of unknown function; possible role in DNA metabolism and/or in genome stability; expression is heat-inducible<ref name='S000043176'>Perkins EL, et al. (1999) Yeast and human genes that affect the Escherichia coli SOS response. Proc Natl Acad Sci U S A 96(5):2204-9 {{SGDpaper|S000043176}} PMID 10051619</ref><ref name='S000114240'>Sakaki K, et al. (2003) Response of genes associated with mitochondrial function to mild heat stress in yeast Saccharomyces cerevisiae. J Biochem 134(3):373-84 |
| − | {{SGDpaper| | + | {{SGDpaper|S000114240}} PMID 14561723</ref> |
<br> | <br> | ||
<br> | <br> | ||
| Line 30: | Line 30: | ||
{{CommentaryHelp}} | {{CommentaryHelp}} | ||
| + | {{UW-Stout}} | ||
| + | ===[[UW-Stout/Heat Shock FA21]]=== | ||
| + | [[Image: Heat Shock.jpeg|center|thumb|400px]] | ||
| + | YPRO78C (D) | ||
| + | *The trials showed very consistent results when heat shocked. The growth rate was consistent but just less clusters than the control plate. | ||
| + | ===[[UW-Stout/Nitrogen_Starvation_FA21| Nitrogen Starvation]]=== | ||
| + | [[Image:YPR078C_Nitrogen21.jpg]] | ||
| + | *In the YPR078C gene, there is a slight increase in growth on the plate that does not contain NH₄. Overall, there is no sign that this particular strain did anything significant to the genes; therefore, makes it resistant to the stress it's being put into. | ||
| − | + | ===[[UW-Stout/Bud Scars FA21]]=== | |
| − | + | ||
| − | + | Cells grown with this gene (YPR078C) knocked out had an average of 1.30 bud scars per cell while the wild type (BY-RY4735) had 0.85 bud scars per cell. Of 109 YPR078C cells counted, 142 bud scars were seen. This indicated that the YPR078C gene slows reproduction rate and thus bud scarring in yeast cells. | |
| − | + | ||
| − | + | ===[[UW-Stout/pH_FA21|pH]]=== | |
| − | - | + | |
| + | [[File:pH_knockout.jpg]] | ||
| + | |||
| + | YPR078C | ||
| + | *When this strain was exposed to a citric acid buffer with a pH of 2.6. At first, the yeast growth appeared stunted and then began to slowly trend upward | ||
| + | |||
| + | Protocol [[https://wiki.yeastgenome.org/index.php/UW-Stout/pH_FA21]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ===[[UW-Stout/G418 FA21]]=== | ||
| + | |||
| + | [[File:YPR078C Vs BY4735 G418.png|left|thumb|400px]] | ||
| + | |||
| + | The YPR078C yeast strain doubling time without any stress was 275 minutes which is much slower than the wild type with 182 minutes. When both strains were stressed by G418 they YPR078C doubling time was 257 minutes and the wild type was 178 minutes. (These times are averaged between three trials) | ||
| + | Protocol [[https://wiki.yeastgenome.org/index.php/UW-Stout/G418_FA21]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ===[[UW-Stout/Ethanol FA21]]=== | ||
| + | [[File:Yeast ToutureV2.jpg|400px]] | ||
| + | * The Yeast Strain of YPR078C was affected by the 18.5 % Ethanol it was almost the exact same as the growth curve as the YMR221C growth curve. | ||
| + | [[UW-Stout/Ethanol FA21]] | ||
| + | |||
| + | |||
| + | ===[[UW-Stout/Copper FA21]]=== | ||
| + | |||
| + | [[File:ypr0.jpg]] | ||
| + | The average doubling time for YPR078W unstressed was 296 minutes, while stressed it was 354. From these observations we can conclude that while stressed with our .5 mM CuSO4 solution, the YBR184W strain grew approximately 17% slower. The unstressed version also had a higher overall size. | ||
| Line 51: | Line 99: | ||
<protect> | <protect> | ||
| + | |||
==References== | ==References== | ||
<!-- REFERENCES ARE AUTOMATICALLY GENERATED. PLEASE DON'T EDIT THIS SECTION--> | <!-- REFERENCES ARE AUTOMATICALLY GENERATED. PLEASE DON'T EDIT THIS SECTION--> | ||
{{RefHelp}} | {{RefHelp}} | ||
</protect> | </protect> | ||
Latest revision as of 14:28, 21 December 2021
Share your knowledge...Edit this entry! <protect>
| Systematic name | YPR078C |
| Gene name | |
| Aliases | |
| Feature type | ORF, Uncharacterized |
| Coordinates | Chr XVI:698265..697147 |
| Primary SGDID | S000006282 |
Description of YPR078C: Putative protein of unknown function; possible role in DNA metabolism and/or in genome stability; expression is heat-inducible[1][2]
</protect>
Contents
Community Commentary
About Community Commentary. Please share your knowledge!
This gene is part of the UW-Stout Orphan Gene Project. Learn more here.
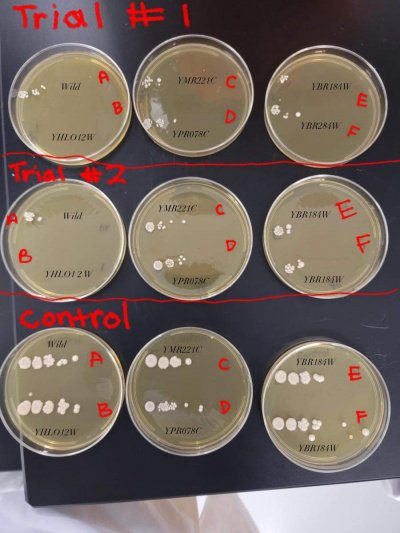
UW-Stout/Heat Shock FA21
YPRO78C (D)
- The trials showed very consistent results when heat shocked. The growth rate was consistent but just less clusters than the control plate.
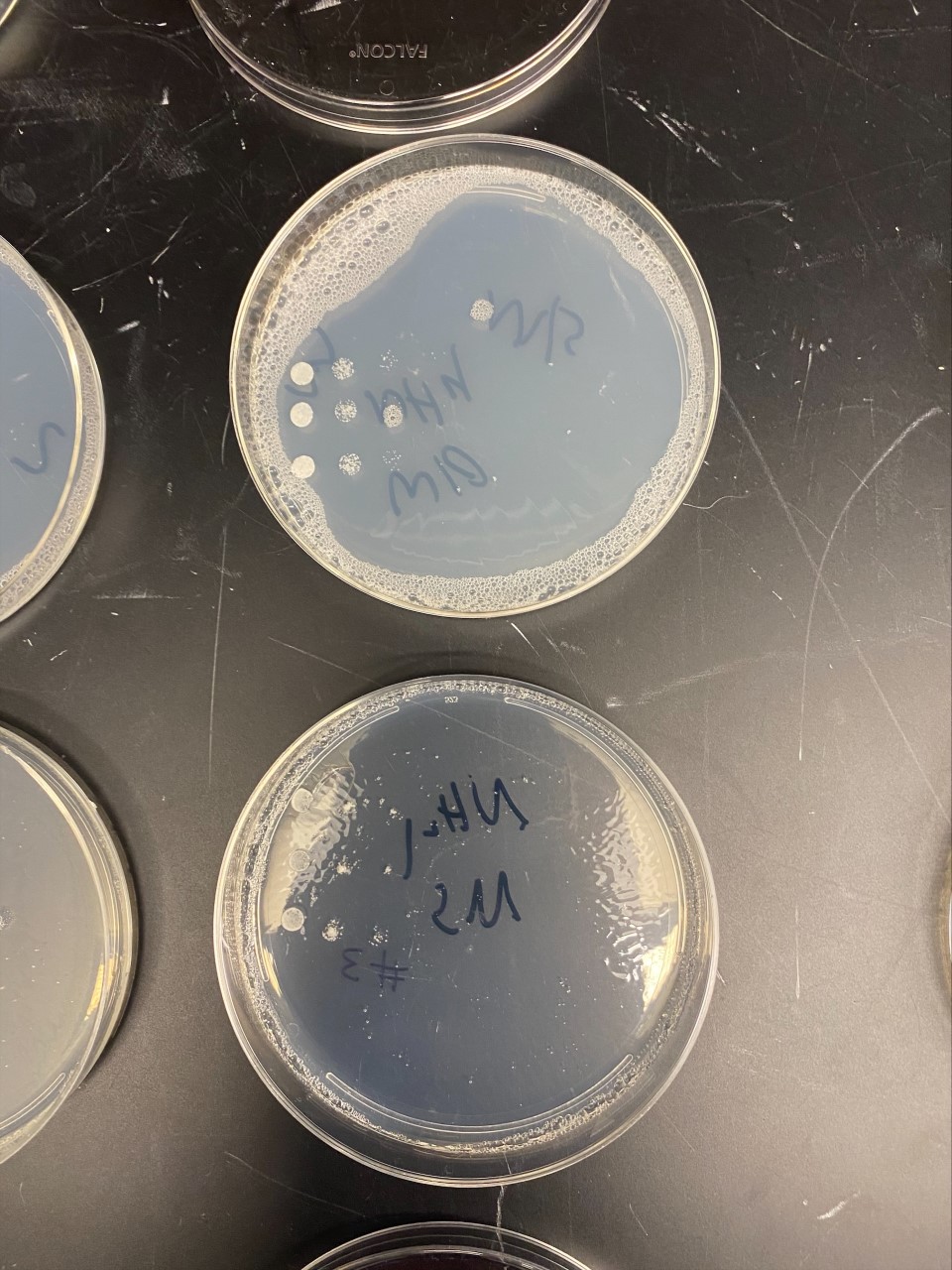
Nitrogen Starvation
- In the YPR078C gene, there is a slight increase in growth on the plate that does not contain NH₄. Overall, there is no sign that this particular strain did anything significant to the genes; therefore, makes it resistant to the stress it's being put into.
UW-Stout/Bud Scars FA21
Cells grown with this gene (YPR078C) knocked out had an average of 1.30 bud scars per cell while the wild type (BY-RY4735) had 0.85 bud scars per cell. Of 109 YPR078C cells counted, 142 bud scars were seen. This indicated that the YPR078C gene slows reproduction rate and thus bud scarring in yeast cells.
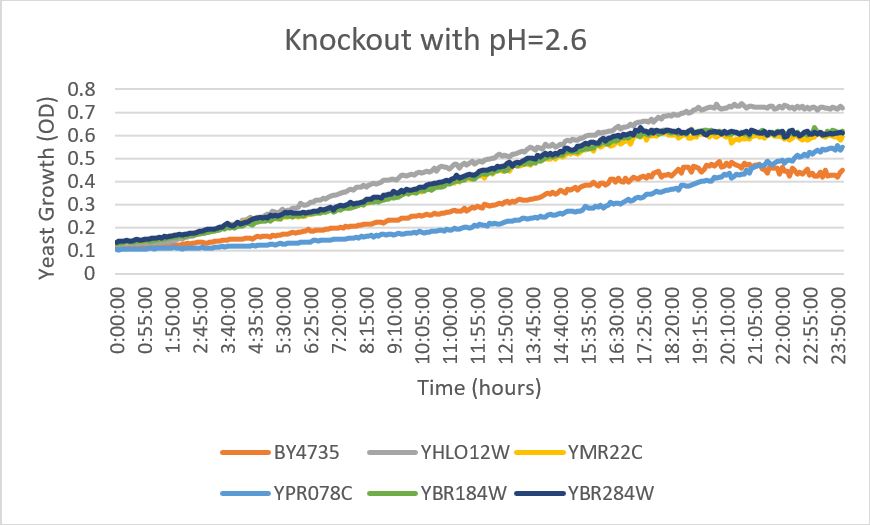
pH
YPR078C
- When this strain was exposed to a citric acid buffer with a pH of 2.6. At first, the yeast growth appeared stunted and then began to slowly trend upward
Protocol [[1]]
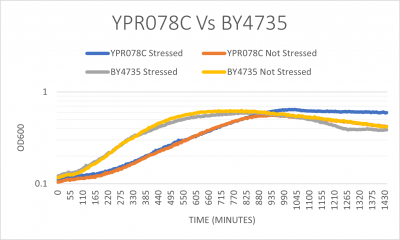
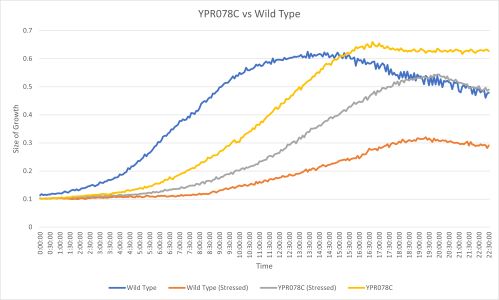
UW-Stout/G418 FA21
The YPR078C yeast strain doubling time without any stress was 275 minutes which is much slower than the wild type with 182 minutes. When both strains were stressed by G418 they YPR078C doubling time was 257 minutes and the wild type was 178 minutes. (These times are averaged between three trials) Protocol [[2]]
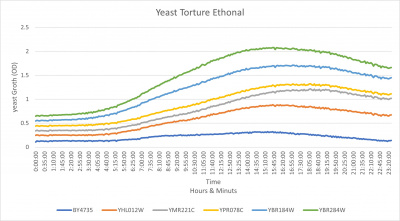
UW-Stout/Ethanol FA21
- The Yeast Strain of YPR078C was affected by the 18.5 % Ethanol it was almost the exact same as the growth curve as the YMR221C growth curve.
UW-Stout/Copper FA21
The average doubling time for YPR078W unstressed was 296 minutes, while stressed it was 354. From these observations we can conclude that while stressed with our .5 mM CuSO4 solution, the YBR184W strain grew approximately 17% slower. The unstressed version also had a higher overall size.
<protect>
References
See Help:References on how to add references
- ↑ Perkins EL, et al. (1999) Yeast and human genes that affect the Escherichia coli SOS response. Proc Natl Acad Sci U S A 96(5):2204-9 SGD PMID 10051619
- ↑ Sakaki K, et al. (2003) Response of genes associated with mitochondrial function to mild heat stress in yeast Saccharomyces cerevisiae. J Biochem 134(3):373-84 SGD PMID 14561723
See Help:Categories on how to add the wiki page for this gene to a Category </protect>