Difference between revisions of "YJR061W"
(→Community Commentary) |
(→Community Commentary) |
||
| Line 56: | Line 56: | ||
The protocol can be found at | The protocol can be found at | ||
===[[UW_Stout/UV_Light_FA19|UV Light Exposure]]=== | ===[[UW_Stout/UV_Light_FA19|UV Light Exposure]]=== | ||
| + | |||
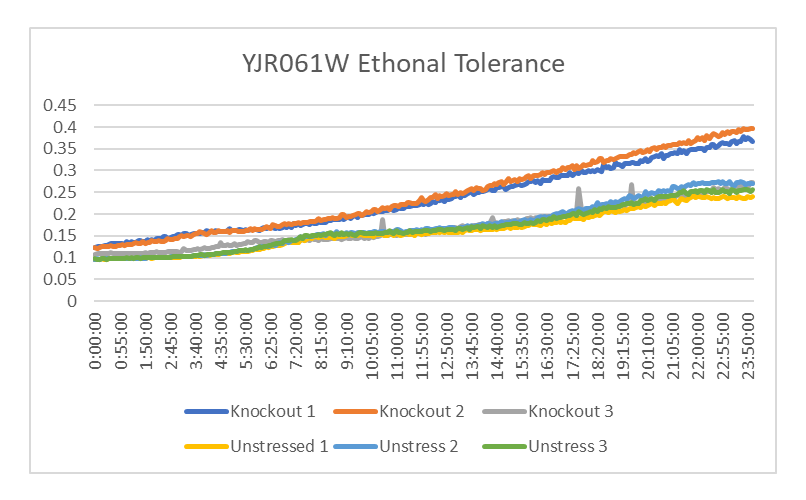
| + | [[Image:YJR061W_ethonal.png]] | ||
<!-- PLEASE ADD Community Commentary ABOVE THIS MESSAGE. See below for an example of community annotation --> | <!-- PLEASE ADD Community Commentary ABOVE THIS MESSAGE. See below for an example of community annotation --> | ||
Revision as of 11:44, 12 December 2019
Share your knowledge...Edit this entry! <protect>
| Systematic name | YJR061W |
| Gene name | |
| Aliases | |
| Feature type | ORF, Uncharacterized |
| Coordinates | Chr X:550511..553318 |
| Primary SGDID | S000003822 |
Description of YJR061W: Putative protein of unknown function; non-essential gene with similarity to Mnn4, a putative membrane protein involved in glycosylation; transcription repressed by Rm101p[1][2]
</protect>
Community Commentary
About Community Commentary. Please share your knowledge!
This gene is part of the UW-Stout Orphan Gene Project. Learn more here.
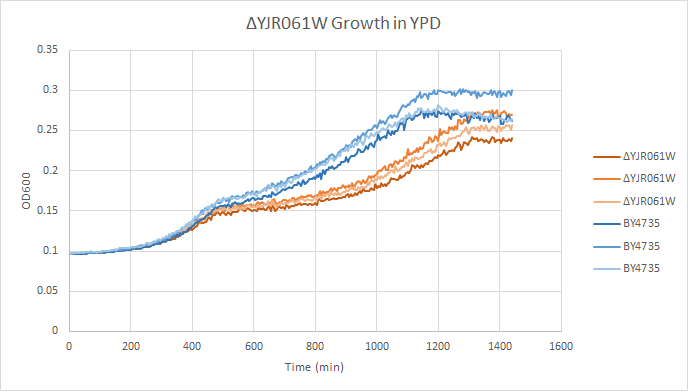
Growth in YPD
In a BY4735 background, knocking out YJR061W seems to have little to no effect on growth rate in log-phase. In this assay, the BY4735 strain's doubling time was 414 minutes, while the YJR061W knock-out strain's doubling time was 436 minutes. (These doubling times are the means of three experiments.)
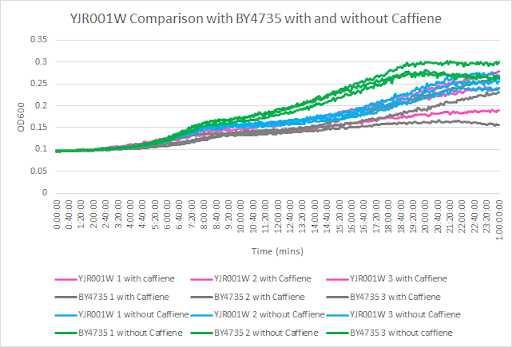
Caffeine and Yeast Cells
This graph shows the growth curve with and without caffeine. It has the knockout strain and wild type yeast cells. The caffeine enhanced the growth in both wild type and knockout strain. The average doubling time between the three experiments for the knockout strand was 824.60 minutes. The caffeine also had a positive effect on the growth curve for the wild type with the average doubling time being 1245.02 minutes. The results were concluded from the wild type and knockout strand was with this amount of caffeine it leads to an increase in growth.
- Note: The graph should be labeled YJK061W
The protocol can be found at
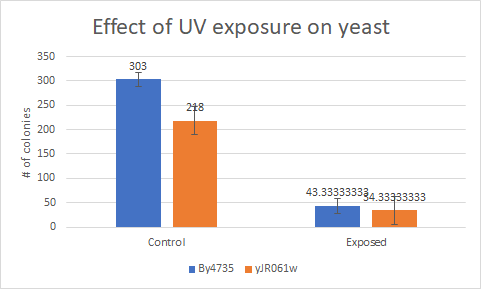
Caffeine
This bar graph showsthat compared to a BY4735 background, Knocking out yJR061w makes yeast a little bit more sensitive to UV light.
The protocol can be found at
UV Light Exposure
<protect>
References
See Help:References on how to add references
- ↑ Conde R, et al. (2003) Screening for new yeast mutants affected in mannosylphosphorylation of cell wall mannoproteins. Yeast 20(14):1189-211 SGD PMID 14587103
- ↑ Lamb TM and Mitchell AP (2003) The transcription factor Rim101p governs ion tolerance and cell differentiation by direct repression of the regulatory genes NRG1 and SMP1 in Saccharomyces cerevisiae. Mol Cell Biol 23(2):677-86 SGD PMID 12509465
See Help:Categories on how to add the wiki page for this gene to a Category </protect>