Difference between revisions of "YBR033W"
(→1% Formamide Yeast Torturehttps://wiki.yeastgenome.org/index.php/UW_Stout/Formamide_FA19) |
|||
| Line 95: | Line 95: | ||
--> | --> | ||
| − | + | ===Zone of Inhibition of 30% [https://wiki.yeastgenome.org/index.php/UW-Stout/Hydrogen_peroxide_FA19 Hydrogen Peroxide] on YKR005C=== | |
Revision as of 14:29, 17 December 2019
Share your knowledge...Edit this entry! <protect>
| Systematic name | YBR033W |
| Gene name | EDS1 |
| Aliases | |
| Feature type | ORF, Uncharacterized |
| Coordinates | Chr II:301944..304703 |
| Primary SGDID | S000000237 |
Description of YBR033W: Putative zinc cluster protein, predicted to be a transcription factor; YBR033W is not an essential gene[1][2]
</protect>
Contents
Community Commentary
About Community Commentary. Please share your knowledge!
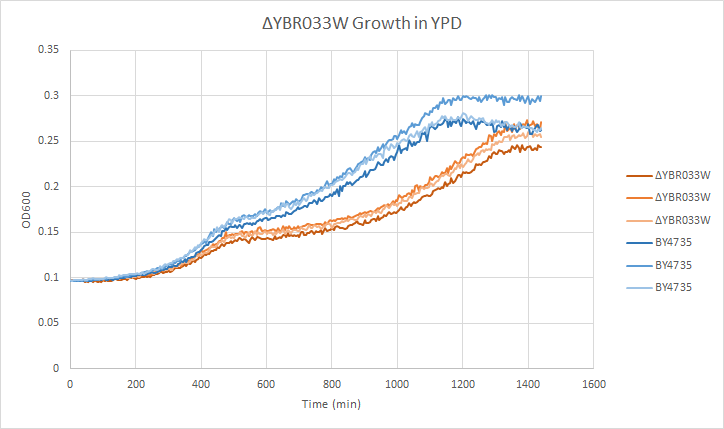
Growth in YPD
In a BY4735 background, knocking out YBR033W seems to have a moderate effect on growth rate in log-phase. In this assay, the BY4735 strain's doubling time was 414 minutes, while the YBR033W knock-out strain's doubling time was 532 minutes. (These doubling times are the means of three experiments.)
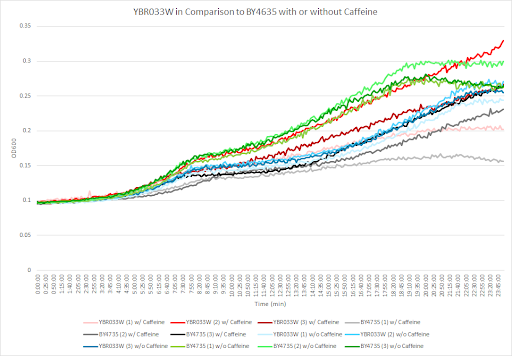
Caffeine and Yeast Cells
This graph shows the growth curve with and without caffeine. It has the knockout strain and wild type yeast cells. The caffeine enhanced the growth in both wild type and knockout strain. The average doubling time between the three experiments for the knockout strand was 708.67 minutes. The caffeine also had a positive effect on the growth curve for the wild type with the average doubling time being 1245.02 minutes. The results were concluded from the wild type and knockout strand was with this amount of caffeine it leads to an increase in growth.
The protocol can be found at
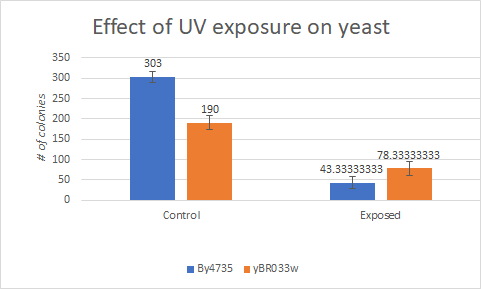
Caffeine
This bar graph show that compared to a BY4735 background, Knocking out yBR033w make yeast less sensitive to UV light exposure
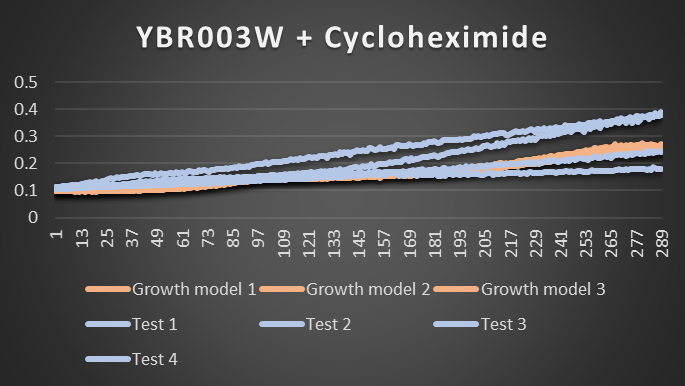
Cycloheximide effecting YBR033W
Growth Models 1-3 are not being stressed. Tests 2-4 are surviving with cycloheximide being added. And Test 1 died which again is different data and weird.
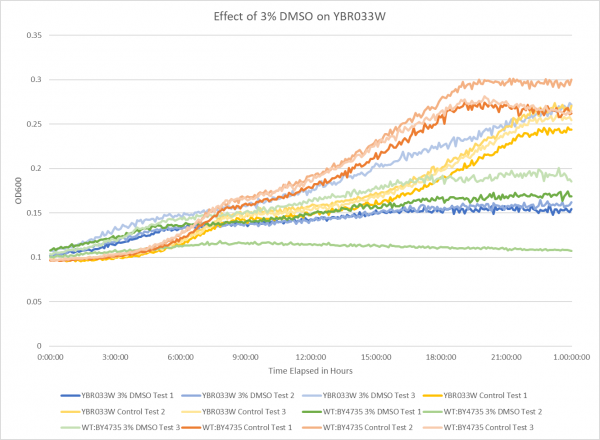
Effect of 3% DMSO on YBR033W
The graph above shows the effect of 3% DMSO on YBR033W. From the data represented the average doubling rate for the YBR033W knockout strain without DMSO was 634 minutes. When DMSO is added the average doubling time increased to 1292 minutes. This shows that 3% DMSO did add stress to this knockout strain. The average doubling time of WT:BY4735 without DMSO was 514 minutes. The average doubling rate for WT:BY4735 with DMSO was 1529 minutes. If we compare the average doubling rate of WT:BY4735 with DMSO to YBR033W with DMSO we can see that without the YBR033W gene the yeast was able to grow more quickly under DMSO induced stress.
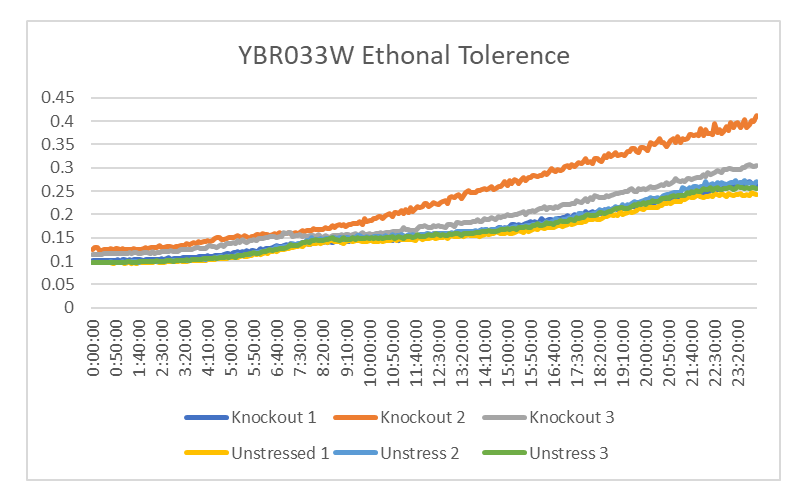
Ethanol Tolerance in YBR033W
This graph shows the growth between YBR033W treated with ethanol and not treated with ethanol. The growth rate of the YBR033W treated with ethanol is 1170.435 minutes. The growth rate of the YBR033W that wasn't treated with ethanol is 1291.754 minutes. The graph shows that the yeast treated with ethanol had a decrease in growth compared to those that weren't.
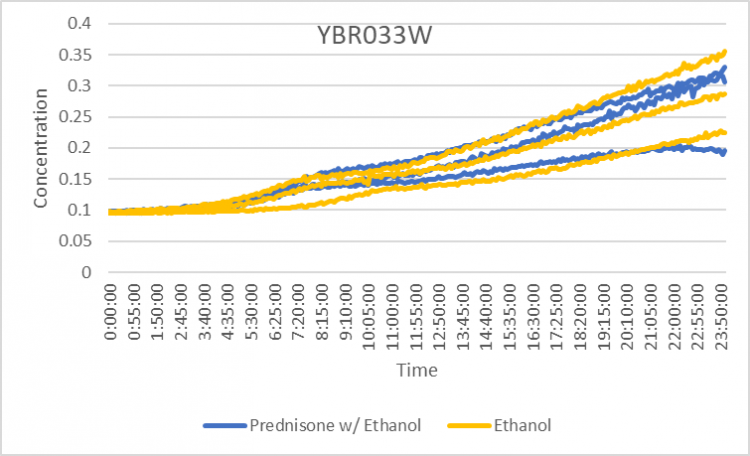
Prednisone Effects on YBR033W
The graph above shows how dilutions P2 (30 µg/ml) and E2 (10% ethanol solution) impacted the growth rate of the knockout yeast strain YBR033W over 24 hours. The doubling rate averages to around 978 minutes for only ethanol. As for the Prednisone with ethanol, the doubling rate averages to around 935 minutes. Overall, ethanol decreased the rate of growth by only about 43 minutes. These results conclude that Prednisone very slightly promotes the growth of yeast cells without the gene YBR033W gene.
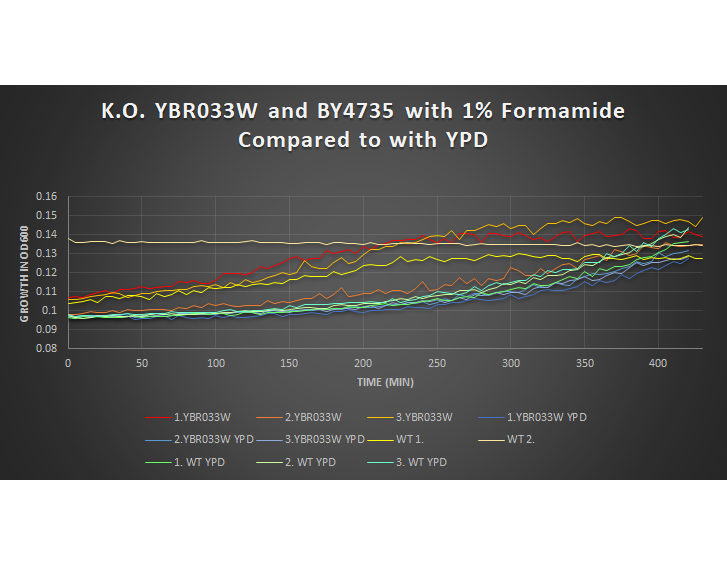
1% Formamide Yeast Torture[[1]]
This graph shows the growth curve of the knockout yeast strain YBR033W and wild type BY4735 in 1% formamide in 3 trials. The average calculated doubling time for YBR033W with 1% formamide was 3,771.50 minutes, while the unstressed strain was 3,181.55 minutes. This concluded that the stressed out strain is slightly sensitive to 1% formamide, and grows as a slower pace in this environment. Our stressed wild types doubling time was 3,459.91 minutes, and the unstressed was 2,468.04 minutes. Again, proving that the addition of 1% formamide constrains growth.
Zone of Inhibition of 30% Hydrogen Peroxide on YKR005C
<protect>
References
See Help:References on how to add references
- ↑ MacPherson S, et al. (2006) A fungal family of transcriptional regulators: the zinc cluster proteins. Microbiol Mol Biol Rev 70(3):583-604 SGD PMID 16959962
- ↑ Yang Y, et al. (2010) Identifying cooperative transcription factors by combining ChIP-chip data and knockout data. Cell Res 20(11):1276-8 SGD PMID 20975739
See Help:Categories on how to add the wiki page for this gene to a Category </protect>